Artifact Removal
A nice artifact detection and removal walkthrough can be found at FieldTrip Artifact Documentation
Visual Artifact Removal
This section addresses visual artifact removal using the FieldTrip toolbox FieldTrip Visual Artifact walkthrough
Load the data and define the trials as per Loading Amplipex data into fieldtrip. Once this has been done we can begin by setting up our cfg (configuration) structure and using the ft_databrowser function
cfg = data_trl.cfg; cfg.channel = 'all'; cfg.trials = 'all'; cfg.latency = 'maxperlength'; cfg.method = 'Summary'; cfg.keepchannel = 'yes'; cfg.metric = 'var'; cfg.alim = []; cfg.coninuous = 'yes'; cfg.viewmode = 'vertical'; % this shows all of the channels vertically cfg = ft_databrowser(cfg,data_trl); data_trl.cfg = cfg; % rewrite the cfg back into the data set
The browser will let you scroll through the trials and use the cursor to highlight a section of the data containing an artifact.
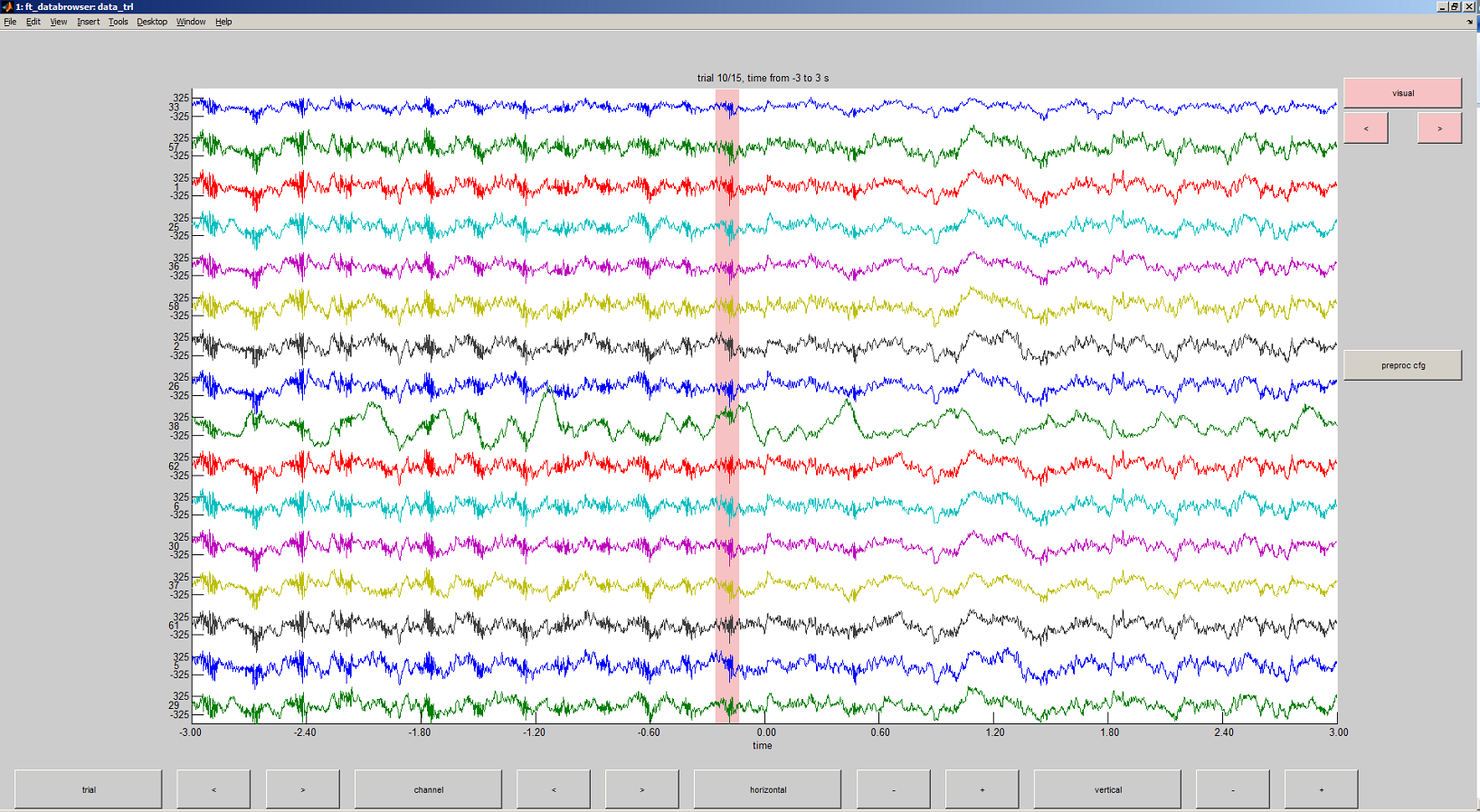
The selected artifact regions are not altered within the data itself, instead the start and end of each artifact are saved as part of the cfg structure: data_trl.cfg.artfctdef.visual.artifact . So now can change these regions to any value of our choice.
%% Mannual Artifact removal art_value = 0; %Value to replace the data within artifact regions artifacts = data_trl.cfg.artfctdef.visual.artifact; trials = data_trl.cfg.trl; for iArt = 1:length(artifacts) for iTrial = 1:length(trials) if trials(iTrial,1)<= artifacts(iArt,1) && trials(iTrial,2)>= artifacts(iArt,2) if artifacts(iArt,1)-trials(iTrial,1) ==0 data_trl.trial{1,iTrial}(1:16,(1:artifacts(iArt,2)-trials(iTrial,1)))=art_value; else data_trl.trial{1,iTrial}(1:16,(artifacts(iArt,1)-trials(iTrial,1):artifacts(iArt,2)-trials(iTrial,1)))=art_value; end end end end
Now we can visualize the data regions that have been replaced to ensure they have been correctly converted.
cfg.viewmode = 'vertical'; % this shows all of the channels vertically ft_databrowser(cfg,data_trl);
From here we can run any further ft analysis.